Collision Cross Sections of Proteins and Their Complexes: A Calibration Framework and Database for Gas-Phase Structural Biology


Traveling‐wave ion mobility mass spectrometry of protein complexes: accurate calibrated collision cross‐sections of human insulin oligomers - Salbo - 2012 - Rapid Communications in Mass Spectrometry - Wiley Online Library

Evaluation of Collision Cross Section Calibrants for Structural Analysis of Lipids by Traveling Wave Ion Mobility-Mass Spectrometry

Collision Cross Sections of Charge-Reduced Proteins and Protein Complexes: A Database for Collision Cross Section Calibration

Figure 1 from Conformational Ordering of Biomolecules in the Gas Phase: Nitrogen Collision Cross Sections Measured on a Prototype High Resolution Drift Tube Ion Mobility-Mass Spectrometer
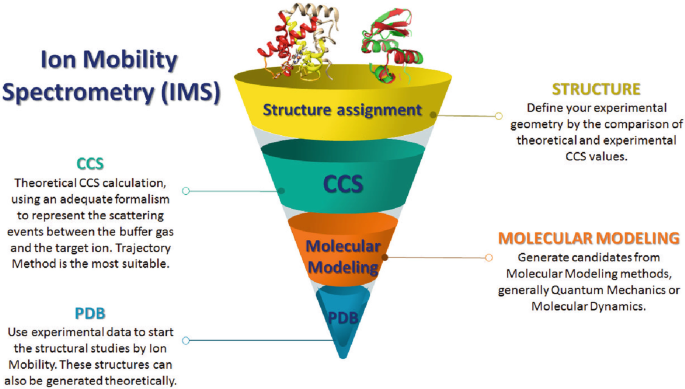
Collision Cross Section Calculations Using HPCCS

Collision Cross-Section Calibration Strategy for Lipid Measurements in SLIM-Based High-Resolution Ion Mobility

Conformational Ordering of Biomolecules in the Gas Phase: Nitrogen Collision Cross Sections Measured on a Prototype High Resolution Drift Tube Ion Mobility-Mass Spectrometer

PDF] Resource Collision Cross Sections f or Structural Proteomics Graphical Abstract Highlights

Collision Cross Sections of Proteins and Their Complexes: A Calibration Framework and Database for Gas-Phase

Mass spectrometry-enabled structural biology of membrane proteins - ScienceDirect

Molecules, Free Full-Text

Collision Cross Sections of Charge-Reduced Proteins and Protein Complexes: A Database for Collision Cross Section Calibration

Ion Mobility Mass Spectrometry (IM-MS) for Structural Biology: Insights Gained by Measuring Mass, Charge, and Collision Cross Section

Ion Mobility Mass Spectrometry (IM-MS) for Structural Biology: Insights Gained by Measuring Mass, Charge, and Collision Cross Section. - Abstract - Europe PMC

Collision Cross Sections of Proteins and Their Complexes: A Calibration Framework and Database for Gas-Phase Structural Biology